The lab works on robust, identifiable and interpretable machine learning tools for scientific inference. We work on both core machine learning topics and the application of machine learning algorithms for scientific inference in the life sciences. Our research focus is on:
- Representation learning, especially using contrastive learning and generative pre-training
- Hierarchical representation learning
- Identifiability and robustness of machine learning algorithms
- (Nonlinear) dynamical system modeling
The following is a list of selected pre-prints and publications published by lab members (including work done at previous institutions). For a full list, please see Google Scholar.
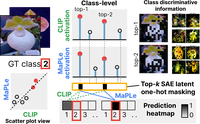
Sparse Autoencoders Reveal Selective Remapping of Visual Concepts During Adaptation
PatchSAE is a framework to extract interpretable concepts and their spatial attributions in vision transformers.
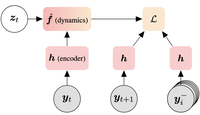
Self-supervised contrastive learning performs non-linear system identification
DynCL is a framework for non-linear system identification using contrastive learning.
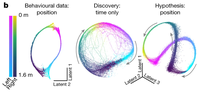
Learnable Latent Embeddings for Joint Behavioral and Neural Analysis
CEBRA is a contrastive learning algorithm for relating behavior and neural activity.
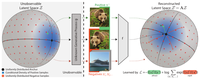
Contrastive Learning Inverts the Data Generating Process
Contrastive learning with the InfoNCE objective can recover the ground truth latent factors underlying the data.
A Primer on Motion Capture with Deep Learning: Principles, Pitfalls and Perspectives
We discuss principles of pose estimation algorithms and highlight their potential and pitfalls for experimentalists.
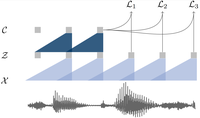
vq-wav2vec: Self-Supervised Learning of Discrete Speech Representations
Learning discrete representations of speech yields state-of-the-art recognition performance on TIMIT and WSJ.
Iron-Sequestering Nanocompartments as Multiplexed Electron Microscopy Gene Reporters
Multicolored gene reporters for light microscopy are indispensable for biomedical research, but equivalent genetic tools for electron …
wav2vec: Unsupervised Pre-training for Speech Recognition
Contrastive pre-training on speech at scale reduces the need for labeled data.
Multi-Task Generalization and Adaptation between Noisy Digit Datasets: An Empirical Study
Transfer learning for adaptation to new tasks is usually performed by either fine-tuning all model parameters or parameters in the …
Salad: A Toolbox for Semi-supervised Adaptive Learning Across Domains
We introduce salad, an open source toolbox that provides a unified implementation of state-of-the-art methods for transfer learning, …
NeuBtracker — imaging neurobehavioral dynamics in freely behaving fish
A long-standing objective in neuroscience has been to image distributed neuronal activity in freely behaving animals. Here we introduce …
Context-based Normalization of Histological Stains using Deep Convolutional Features
While human observers are able to cope with variations in color and appearance of histological stains, digital pathology algorithms …